New Study Reveals Why Bacteria Fail to Stop “Superbug” Plasmids
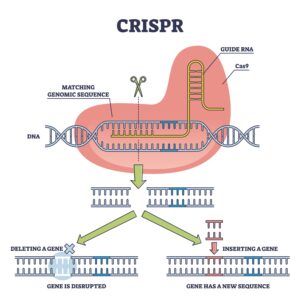
In the microscopic world of bacteria, a silent and deadly arms race is constantly unfolding. On one side are plasmids—rogue loops of DNA that jump from one cell to another, often carrying the instructions for antibiotic resistance. On the other side is CRISPR, the famous “genetic scissors” that bacteria use as an immune system to chop up these invaders.
But if CRISPR is so effective, why does antibiotic resistance still spread so quickly? A groundbreaking study in PNAS has finally provided the answer: it’s all about the kinetics—the speed and timing of molecular reactions.
The “Tug-of-War” in a Single Cell
For years, scientists studied CRISPR in a vacuum, looking at how it works in a single test tube. However, this team used high-resolution “time-lapse” microscopy to watch the battle happen in real-time inside living E. coli cells.
They discovered that CRISPR isn’t a simple “on/off” switch. Instead, it’s a high-stakes race. When a plasmid enters a cell, the CRISPR system has a very narrow window of time to find and destroy it. If the CRISPR molecules move too slowly, or if there aren’t enough of them, the plasmid can “set up shop,” replicating itself and even moving on to the next cell before the defense system can react.
The “Addiction” Factor
The researchers found a devious trick used by plasmids to win this race: Plasmid Addiction. Some plasmids carry “toxin-antitoxin” systems. If the CRISPR system successfully destroys the plasmid, the cell is left with a lingering toxin that kills the host.
In these cases, the CRISPR “defense” actually becomes a death sentence for the bacterium. This creates a fascinating paradox: the bacteria that are best at defending themselves might be the first to die, allowing the “weaker” bacteria—who have accepted the plasmid—to take over the colony.
While this research happens at the scale of molecules, its impact is human-sized.
-
Stopping Superbugs: By understanding the “speed limit” of CRISPR, scientists can design better ways to stop the spread of antibiotic resistance. If we can slow down the plasmid’s “setup” time or speed up the CRISPR “search” time, we could potentially “de-program” resistance from a bacterial population.
-
Precision Medicine: This study allows us to predict how a bacterial community will evolve. Instead of guessing if a treatment will work, doctors might one day use “agent-based models” (the same tech used in this study) to simulate how a specific infection will react to different genetic interventions.
-
Synthetic Biology: For industries that use bacteria to create medicines like insulin, keeping “good” plasmids in and “bad” ones out is vital. This research provides a “user manual” for fine-tuning those genetic defenses to ensure industrial vats of bacteria remain productive and healthy.
The Big Picture
The study proves that we cannot understand the “forest” (the bacterial population) without understanding the “trees” (the molecular kinetics). By measuring how fast individual proteins move and bind, we can now predict whether a whole city of bacteria will succumb to a new genetic invader or stand its ground. It is a masterclass in how the smallest movements in nature dictate the grandest outcomes of evolution.


